Labnetin vieraskynäblogi
As more and more diseases are linked to epigenetic abnormalities, there has been an exponential growth in research focused on the association between epigenetics and disease. As such, laboratories new to the areas of epigenetics and gene regulation research are experiencing a growing need for tools to facilitate the innovation of research into these areas.
Active Motif is the industry leader in developing and delivering innovative tools to enable epigenetics and gene regulation research. Their staff of highly qualified scientists work in close collaboration with world-class researchers to continually develop new and innovative tools to study chromatin, histones, DNA methylation, non-coding RNAs and transcription biology.
Epigenetics and disease
The epigenetics state of the cell is controlled by the activity of proteins that add and remove small chemical modifications to histones and to DNA. Aberrant epigenetic regulation can lead to changes in gene expression and human disease (table 1 & 2).
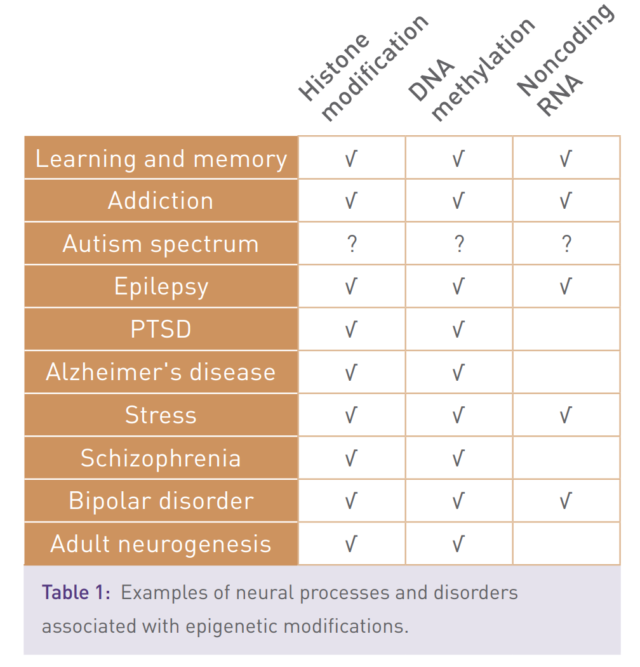
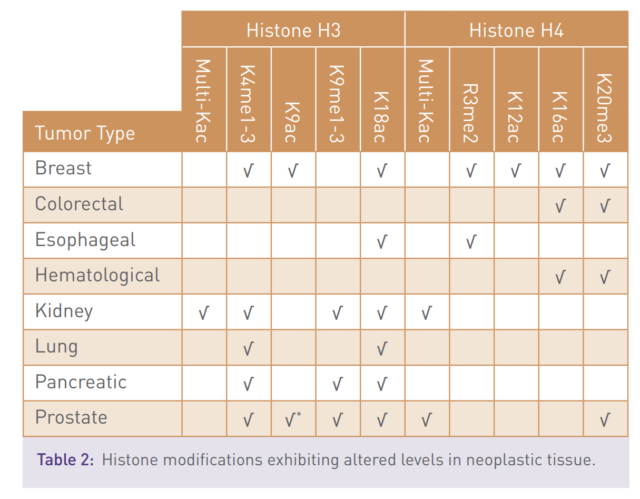
The link between epigenetics and disease has been substantiated through the identification of mutations in, or altered expression of, epigenetic regulator proteins that are associated with a multitude of pathological states and poor disease prognosis.
These mutations have been identified in all three major classes of epigenetic proteins: the Writers (enzymes that deposit modifications), the Erasers (enzymes that remove modifications) and the Readers (proteins that recognize and bind epigenetic modifications). Recent drug development strategies that target these enzymes have proved highly successful, resulting in four FDA approved cancer drugs, with many more promising leads on the horizon.
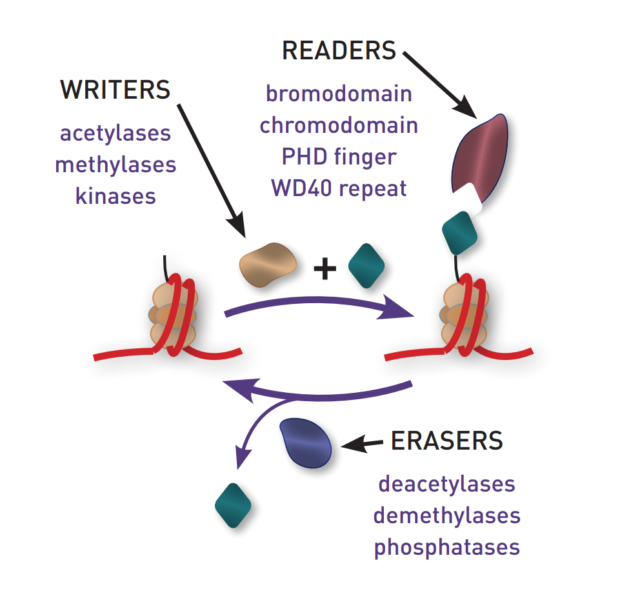
The epigenetic tool kit
The development of methods to perform chromatin immunoprecipitation (ChIP) and analyses of DNA methylation, histones and gene regulation, along with Next-Generation sequencing tools, is allowing researchers to get a better picture of the genome-wide changes associated with cancer and other diseases. Active Motif provides products that utilize familiar techniques and reagents, or simplify technologies, to enable even a novice to perform epigenetic studies. These include:
- Antibodies to discriminate between the multiple post-translational modifications on the histone tails and to detect methylated DNA
- ChIP and ChIP-Seq techniques to enable scientists to link specific states of chromatin to individual gene loci in a cell to understand how genes are regulated
- ELISA-based methods for analysis of global changes in levels of intercellular markers, such as histone modifications, DNA methylation and transcription factor binding
- Luciferase reporter assays to study the impact of promoter regulation and miRNA-3’UTR interactions on transcriptional and post-transcriptional regulation of genes
Examples of methods to map disease-specific epigenetic variances:
Uncover epigenetic variances in archived tissue
Large collections of archived formalin-fixed, paraffin embedded (FFPE) tissue are available that harbor invaluable information about the variances between normal and diseased tissue states. However, performing CHIP has not been possible on FFPE samples because of the difficulty in extracting high-quality chromatin from very limited and degraded material. Active Motif’s ChIP FFPE assay is the first-of-its-kind to enable extraction and ChIP analysis of chromatin from FFPE samples for Next-Generation sequencing.
Evaluate chromatin changes in primary cells
Because of the importance of PBMCs in cancer and immunology/ infectious disease research, a robust method to extract quality chromatin from these cells for ChIP and ChIP-Seq analysis is highly warranted. PBMCs make it difficult to extract quality chromatin because they are highly resistant to lysis under conditions normally suitable for other cell types. Active Motif scientists have developed an optimized chromatin preparation method that yields high-quality chromatin from primary cells for ChIP analysis and Next-Generation sequencing. Active Motif’s ChIP-IT PBMC Kit is the only ChIP kit available to enable ChIP to be performed on difficult-to-lyse peripheral blood mononuclear cells, or PBMCs, including lymphocytes (T cells, B cells & NK cells) and monocytes.
Measure global changes in the epigenome – generate DNA methylation & histone profiles of your cell models
Because changes in global methylation are hallmark of many human diseases, including cancer, simpler methods than HPLC or bisulfite sequencing are warranted for correlative studies that analyze variances in genome-wide DNA methylation status. Active Motif’s Global DNA Methylation – LINE-1 assay uses a unique hybridization approach that quantitates 5-methylcytosine (5-mC) levels at LINE-1 repeats as surrogate measure of global methylation. This unique hybridization approach offers better specificity and reproducibility than other available methods that utilize passive adsorption.
The addition or removal of modifications such as phospho-, methyl- and acetyl- functional groups to histones have a profound effect on the regulation of transcription, chromosome packaging and DNA damage repair. Screening extracts for specific histone modifications is a simple way to access cell health and compound effects. Active Motif’s comprehensive selection of Histone Modification ELISAs provides a simple solution for screening changes in histone H3 modification levels from purified core histones or histones isolated by acid extraction.
Did you get interested?
You can learn more about these tools from the Active Motif’s Tools for Disease Research brochure. There is also information how to:
- measure transcription factor activity
- identify biological responses to compound treatments
- study gene regulation with reporter assays
- measure miRNA function and validate miRNA targets
- study immune function and inflammatory response
and much more…
Koonnut: Heidi Bondén / Labnet



